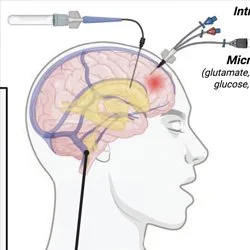
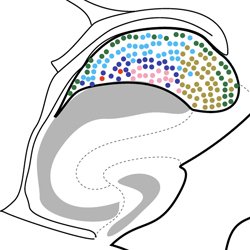
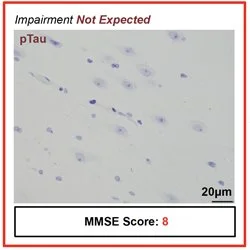
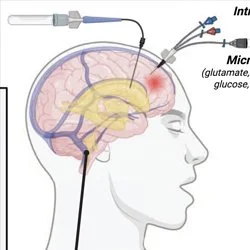
What does a memory “look like” in the brain? How do memory and cognition go awry in brain disorder?
These are the questions we seek to answer in our multidisciplinary, highly collaborative lab. Our research combines cutting-edge experimental techniques with computation, engineering, and mathematics to study memory and how it changes in brain disorder and disease. Our primary research applications are centred around Alzheimer’s disease, anxiety and fear memory, and epilepsy, with our lab also having far-reaching interests including recognition memory, traumatic brain injury, autism spectrum disorder, and schizophrenia. We gratefully acknowledge the supporters of our research for making this work possible.
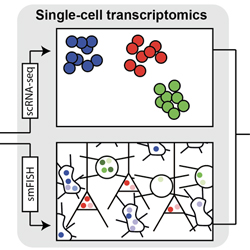
MOLECULAR
Goal: identification of cell types and associated key molecules
Molecular techniques:
Single-cell RNA-seq
Single-cell spatial transcriptomics
CELLULAR
Goal: validate and causally interpret cell types and molecules
Cellular techniques:
Patch-clamp electrophysiology
Cell-type-specific gene knockouts
CIRCUIT
Goal: map molecules and cell types onto circuits and develop circuit-specific access
Circuit techniques:
Virus-mediated anatomical mapping
Virus-mediated optogenetic and chemogenetic manipulations
BEHAVIOURAL
Goal: interpret behavior with molecular, cell-type, and circuit specificity
Behavioural techniques:
Large-scale Neuropixels electrophysiology
1-photon calcium imaging during freely moving behaviour
2=photon calcium imaging during head-fixed behaviour
Optogenetic manipulation during behaviour
CLINICAL
Goal: translate and reverse-translate results between rodents and humans
Clinical techniques and applications:
Tissue biospecimens from brain disorder and disease
Clinicopathological correlations
Acute and organotypic slice culture of living human brain tissue
2-photon imaging of living human brain tissue following drug intervention
Recent News
Summer 2026: Mark is elected as the Vice President (and President-elect) of the Canadian Association for Neuroscience. Catie Futhey receives the Necia Laura May Elvin Memorial Prize in Schizophrenia Research. Adrienne Kinman receives a CAN-CIHR-INMHA Brain Star Award for her 2025 publication on ovoid neurons. Tomos Salathiel receives an Alzheimer’s Society of British Columbia and Yukon Doctoral Scholar award. Postdoctoral researcher Mehwish Anwer is headed to the University of Lethbridge as a tenure-track faculty member. In a collaboration with lead investigator Dr. Mypinder Sekhon (UBC Critical Care Medicine), the lab receives a VCHRI Innovation and Translational Research Award funded by the VGH & UBC Hospital Foundation and Organ Donation and Transplant Research Foundation of BC.
Spring 2026: Catie Futhey receives a Canadian Institutes of Health Research Doctoral Scholarship. Julia Groening receives a Canada Graduate Research Scholarship - Master’s. Tom van der Sloot receives an NSERC Undergraduate Student Research Award. Muskan Poddar receives an Undergraduate Program in Neuroscience Science Undergraduate Research Experience award. Cindy Zhang receives a School of Biomedical Engineering Synergy Award as well as a Faculty of Science Enhanced Science Undergraduate Research Experience award. Work on clinicopathologic subtypes of schizophrenia, led by MD/PhD student Catie Futhey in collaboration with Veronica Hirsch Reinshagen, is published in Translational Psychiatry.
Winter 2026: A review on subiculum cell types led by Adrienne Kinman is published in Trends in Neurosciences. Research led by Regan Campbell on specialized cellular computations mediated by covarying morphological-hodological properties is published in Progress in Neurobiology. Our review on applications of -omics technology in critical care, co-authored with Jordan Bird and Myp Sekhon, is published. Tom van der Sloot is named a Wesbrook Scholar (one of 20 scholars named across all of UBC undergraduate, MD, and JD students), as well as an HSBC Emerging Leader Scholar and a Thelma Sharp Cook Scholar. Tom is also named as a Canada Co-op Student of the Year. Margarita Kapustina receives the UBC Department of Statistics Award in Data Science. The lab receives a collaborative CIHR Project Grant, led by Dr. Mypinder Sekhon, to study brain function during post-mortem extra-corporeal membrane oxygenation.
Fall 2025: Catie Futhey is named a Li Tze Fong Memorial Fellow at UBC. Julia Groening receives an Experimental Medicine Graduate Student Entrance Award. Larissa Kraus receives the Bluma Tischler Postdoctoral Fellowship. Work on computational analysis tools for mFISH analysis, led by Kaitlin Sullivan, is published in STAR Protocols. Work on analysis of Layer 6b neurons across the cortex, led by Margarita Kapustina, is published in Progress in Neurobiology.
Summer 2025: Adrienne Kinman defends her PhD, and is highlighted by the Globe & Mail as an emerging star of Canadian science! Aoi Ichiyama joins the lab as a postdoc. Research examining neural reorganization following traumatic brain injury, done in collaboration with the Wellington lab, is published in Cell Reports. A study led by the Plemel lab examining microglia proliferation, for which the our lab contributed to, is published in Development. Margo Kapustina is named a UBC Killam Doctoral Fellow. Mathias Delhaye receives a Graduate Student Rising Star in Neuroscience Award. Tom van der Sloot wins the Glenn and Annetta Agnew Science Co-op Award, the Undergraduate Program in Neuroscience Leadership Award, the Charles and Jane Banks Scholarship, and is a runner up for the Undergraduate Program in Neuroscience Research Award. Brianna Bristow receives the Friedman Award for Scholars in Health. Mark leads a ~$5M UBC team grant to integrate multimodal data across -omics and functional imaging.
Spring 2025: Dr. Kaitlin Sullivan defends her PhD, and Kate Dunne completes her BSc honours thesis! Julia Groening joins the lab for her MSc. Brianna Bristow receives the Laura G. Jasch Memorial Prize. Kate Dunne and Ming Zhang both receive Science Undergraduate Research Experience awards. Kate Dunne also receives the Edgar C. Black Memorial Prize and the Kurt Henze Memorial Prize. Riya Gandhi receives a School of Biomedical Engineering Synergy Award. Regan Campbell and Margarita Kapustina each receive NSERC Doctoral Scholar Awards. Ryan Ripsman receives the Alzheimer’s Society of Canada Doctoral Scholar award. Mark receives the Canadian Association for Neuroscience New Investigator Award. The Cembrowski lab is now on Instagram.
Winter 2025: Work led by Adrienne Kinman, discovering and interpreting an atypical excitatory neuron in the subiculum that governs object memory, is published in Nature Communications and has been highlighted by a variety of news sources around the world, as well as a viral social media post. Adi Swaro is named a Wesbrook Scholar (one of 20 scholars named across all of UBC undergraduate, MD, and JD students), as well as an HSBC Emerging Leader Scholar. The lab receives a CIHR Project Grant to study causal cell types in epilepsy.
Fall 2024: Daemon Cline is awarded a CELL Fellowship, Regan Campbell is awarded a UBC Four-Year Fellowship, Margarita Kapustina is awarded a UBC Kruger Graduate Fellowship, Adi Swaro is awarded the Neuroscience Program Third-Year Research Award and is a runner up for the Neuroscience Program Third-Year Leadership Award. Mark is awarded an Alzheimer Society Research Program New Investigator grant to study the cellular causes of Alzheimer’s disease from the Alzheimer’s Society of Canada and Brain Canada, and a Future Leader of Brain Canada Momentum Grant from Brain Canada to study causal cell types in epilepsy. The lab is featured in Pathways.
Summer 2024: Shalini Iyer, Daemon Cline, and Margarita Kapustina each receive Endowment Awards from the Djavad Mowafaghian Centre for Brain Health. Adi Swaro receives a Summer Student Research Program award and a Science Undergraduate Research Experience Award. Larissa Kraus is awarded a CIHR postdoctoral fellowship, Catie Futhey is awarded a CIHR Canada Graduate Scholarship - Masters, and Brianna Bristow is awarded an NSERC Canada Graduate Scholar-Doctoral Award. The Cembrowski Laboratory is awarded an NSERC Discovery Grant to study Layer 6b of the neocortex.
Spring 2024: Ming Zhang is awarded an SBME Synergy Award, Kate Dunne is awarded an NSERC Undergraduate Student Research Award, and Kaitlin Sullivan is awarded the Laura G. Jasch Memorial award. Work from the lab on the anterior thalamic nuclei is published in Cell Reports. Sarah Erwin, MSc, successfully defends her master’s thesis. Brianna Bristow is awarded the Dorothy May Ladner Memorial Fellowship from the UBC Faculty of Medicine.
Fall 2023: Shalini Iyer is interviewed on Global News for her leadership and advocacy in the Support Our Science movement. Kaitlin Sullivan is awarded a Friedman Award for Scholars in Health Fellowship.
Summer 2023: Adi Swaro wins the Outstanding Second-Year Student Leadership in Neuroscience Award, and receives second-place for his student presentation at the 2023 SBME Synergy Research Day. Brianna Bristow and Regan Campbell are awarded DMCBH General Awards, and Catie Futhey is awarded the Schizophrenia Endowment Award. Shalini Iyer is awarded a Postgraduate Scholarship - Doctoral award and a UBC 4-Year Fellowship.
Spring 2023: The lab, in collaboration with clinical and engineering colleagues, is awarded a New Frontiers in Research Fund grant to study the mechanisms of epilepsy in the human brain. Adi Swaro and Cate Futhey receive SBME Synergy Awards for their summer research. Brianna Bristow and Margarita Kapustina are each awarded Canada Graduate Scholarships - Masters from CIHR. Margarita is also awarded an NSERC Undergraduate Student Research Award. Kaitlin Sullivan is awarded the Department of Statistics Award in Data Science. Derek Merryweather receives both the Faculty-nominated and Student-nominated Best Presentation Award at the CPS Departmental Retreat, and Larissa Kraus scores 2nd place for her poster. Lab alumnus Jennifer Tsai receives both the University of Waterloo and Ontario province co-op student of the year awards for her research in the lab.
People
Principal investigator
Mark S. Cembrowski, MS, PhD Short CV (March 2026) | mark.cembrowski@ubc.ca
Primary affiliation: Associate Professor (with Tenure), Department of Cellular and Physiological Sciences, Faculty of Medicine, University of British Columbia
Other affiliations:
In 2019, Mark started as an Assistant Professor in the Department of Cellular and Physiological Sciences at the University of British Columbia and an Investigator with the Djavad Mowafaghian Centre for Brain Health, and was promoted to Associate Professor (with tenure) in 2024. His research interests are in applying technological, computational, and mathematical approaches to understand healthy brain function, especially learning and memory, as well as brain changes that emerge in disorder and disease. He has received a variety of recognitions for his research, including being named a Next Generation Leader by the Allen Institute, a Future Leader of Canadian Brain Research by the Brain Canada Foundation, a Scholar with Michael Smith Health Research British Columbia, an Alzheimer’s Young Investigator by the Alzheimer’s Society of Canada, the Krieg Cortical Scholar in the 2020 worldwide competition, and the New Investigator Award from the Canadian Association for Neuroscience in the 2025 national competition.
Mark received his BSc in Mathematics from the University of British Columbia (2007), where he conducted research examining complex oscillatory behaviour of a sixth-order nonlinear partial differential equation. He later received his MS (2008) and PhD (2011) in Applied Mathematics from Northwestern University, where he combined computational modeling and patch-clamp electrophysiology to study retinal processing as a joint student between William Kath, Hermann Riecke, and Joshua Singer. As a postdoc, Mark worked in the laboratory of Nelson Spruston at the Janelia Research Campus of the Howard Hughes Medical Institute. His research combined computational modeling, big data analysis, transcriptomics, electrophysiology, viral circuit mapping, and animal behaviour to study the role of cell types in memory. During his training Mark received a National Science Foundation Graduate Research Fellowship, a Royal E. Cabell Fellowship, a Northwestern University Scholarship, and an Natural Sciences and Engineering Research Council of Canada Postgraduate Scholarship.
In his spare time, Mark practises and teaches yoga, handstands, and calisthenics.
Lab members
Larissa Kraus, PhD (Epilepsy Working Group) CV | Google Scholar | larissa.kraus@ubc.ca
CIHR Postdoctoral Fellow | Walter Benjamin Programme Postdoctoral Fellow | Bluma Tischler Postdoctoral Fellow
Larissa’s research focuses on identifying subpopulations of cells in healthy and diseased human brain tissue. In this way, she hopes to identify novel mechanisms to treat neurological diseases, such as epilepsy. Before joining the Cembrowski lab and moving to Vancouver, Larissa completed her PhD in Berlin, Germany (2020).
Aoi Ichiyama, PhD (Mood Disorders Working Group) CV
Postdoctoral Fellow
Aoi completed her PhD in Neuroscience at the University of Western Ontario, where she studied the neural circuits that regulate the stress response. In the Cembrowski lab, she will investigate how the activity of novel hippocampal cell types contributes to behaviour and how their dysregulation manifests in diseases and disorders.
Nick Vyleta, PhD (Healthy Cognition Working Group)
Research Associate
Nick is interested in neuronal physiology and circuit function, and has done extensive work studying synaptic plasticity and network dynamics in hippocampal memory circuits using paired electrophysiology recordings from brain slices. Nick received his PhD from Oregon Health and Science University in Portland, Oregon and his postdoctoral work includes a fellowship at the Institute for Science and Technology, Austria in Vienna before working on adult-neurogenesis at UBC.
Rachel Boyd, PhD (Mood Disorders Working Group) CV
Postdoctoral Fellow
Rachel is an incoming postdoctoral fellow interested in understanding the cellular and molecular mechanisms underlying brain dysfunction in neuropsychiatric and neurodegenerative diseases. Rachel received her BSc in Genetics at the University of Western Ontario, and recently completed her PhD in Human Genetics and Genomics at the Johns Hopkins University School of Medicine, where she studied gene regulatory mechanisms underlying Parkinson's Disease risk.
Mehwish Anwer, PhD (Alzheimer’s Working Group) CV | Google Scholar
CIHR Postdoctoral Fellow | Michael Smith Health Research BC Postdoctoral Fellow | Bluma Tischler Postdoctoral Fellow Postdoctoral Fellow
Mehwish’s research focuses on the spatial and molecular landscape of brain injury and associated dementia including Alzheimer’s Disease. Her work integrates spatial transcriptomics, molecular neuropathology, and human tissue analysis to understand how interactions among neurons, glia, and vascular cells shape regional vulnerability after traumatic brain injury. She aims to map neurovascular dysfunction and glial responses across injury and aging to identify targets that promote brain resilience and repair.
Mathias Delhaye (Epilepsy Working Group) CV
PhD Candidate (co-supervised with Ann Marie Craig)
Mathias' research focuses on assessing synaptic composition in hippocampal subregions in relation to epilepsy. For this purpose, he will adapt the Expansion Microscopy to apply it on human samples. Originally from France, Mathias studied at the École Normale Supérieure Paris-Saclay and is currently registered at the Sorbonne University Master of Neuroscience.
Daemon L. Cline (Healthy Cognition Working Group) CV | Google Scholar | dlcline@student.ubc.ca
PhD Candidate | CELL Fellow | NSERC Canada Graduate Scholar- Doctoral | UBC Four-Year Fellow
Daemon's current research focuses on the neuroendocrinology of energy metabolism by studying hormones produced in the periphery that act on energy-regulating centres of the brain. Daemon completed his BSc in Biochemistry at UNBC with a thesis centred on disturbance ecology of streams in Northern BC. He then earned a MSc of Biochemistry at UNBC in the lab of Dr Sarah Gray, where he studied energy-regulating hormones.
Derek Merryweather (Healthy Cognition Working Group) CV | dmerrywe@student.ubc.ca
PhD Candidate
Derek is interested in the intrinsic properties, circuit architecture, and functional output of the subiculum, as it relates to memory and cognition. He received is BS in human physiology from the University of Oregon, after which he worked as a technician for the GENIE Project at the Janelia Research Campus, and he completed his masters at NYU in Dr. Adam Carter's lab.
Shalini Iyer (Healthy Cognition Working Group) CV | siyer13@student.ubc.ca
PhD Candidate | NSERC Postgraduate Scholar - Doctoral | UBC Four-Year Fellow | BC Research Scholar
Shalini is currently completing her MSc in Neuroscience at York University, investigating the role of a bioactive lipid signalling molecule on hippocampal development and the link to autism spectrum disorder (ASD). In the Cembrowski lab, Shalini is interested in investigating developmental abnormalities of specific cell types within the deep subiculum and neocortex using the valproate model of ASD
Regan Campbell (Healthy Cognition Working Group) CV | recampb@student.ubc.ca
PhD Candidate | NSERC Canada Graduate Scholar - Doctoral | UBC Four-Year Fellow | CIHR Canada Graduate Scholar - Master’s
Regan is combining computational modeling and in-vivo multi-photon calcium imaging to identify how neuronal morphology shapes cellular computations, and how these computations operate in-vivo to give rise to cell-type specific functions in the subiculum. She previously completed her H.BSc in Neuroscience and Applied Statistics at the University of Toronto. During this time, she completed her honours thesis in Dr. Rutsuko Ito’s lab, where she investigated the role of the hippocampus in temporal memory and motivational conflict.
Brianna Bristow (Alzheimer’s Working Group) CV | brianna.bristow@ubc.ca
PhD Candidate | Friedman Scholar | NSERC Canada Graduate Scholar - Doctoral | UBC Four-Year Fellow | Dorothy May Ladner Memorial Fellow | CIHR Canada Graduate Scholar - Master’s
Brianna’s research focuses on characterizing a novel cell type exemplifying a unique transcriptomic profile in the cortex. Previously, Brianna completed her BSc in Biochemistry & Molecular Biology at the University of British Columbia. Here, she completed her thesis work in the Ciernia Lab, where she developed functional assays to measure microglial activation in vitro and optimized media conditions to best model the brain microenvironment.
Catie Futhey (Alzheimer’s Working Group) CV | cfuthey@student.ubc.ca
MD-PhD Candidate (co-supervised with Veronica Hirsch-Reinshagen) Li Tze Fong Memorial Fellow | CIHR Canada Graduate Scholar - Doctoral | CIHR Canada Graduate Scholar - Master’s | UBC Four-Year Fellow
Catie Futhey is an MD/PhD candidate co-supervised by Dr. Veronica Hirsch-Reinshagen, interested in the interface between the brain, heart, and immune system, and how sex differences shape this network. Her research uses human postmortem tissue and spatial proteomics to investigate microscopic signatures in the hippocampus linked to disrupted memory and cognition, focusing on synaptic changes, neuroinflammation, and vascular injury in Alzheimer’s disease and schizophrenia. She holds a BSc in Neuroscience from McGill University. In her spare time she enjoys drumming regular-sized drums.
Ryan Ripsman (Alzheimer’s Working Group) CV | rripsman@student.ubc.ca
MD-PhD Student | Alzheimer’s Society of Canada Doctoral Scholar
Ryan is an MD/PhD student at UBC. He is interested in the interactions between brain tumours and the neuronal microenvironment. Ryan completed his undergraduate degree in physics and his master's degree in laboratory medicine and pathobiology at the University of Toronto. For his master's research he worked on integrating multi-omic liquid biopsy technology into a precision oncology program at SickKids Hospital.
Tomos Salathiel (Alzheimer’s Working Group)
MD-PhD Student | Alzheimer’s Society of BC and Yukon Doctoral Scholar
Tomos is an incoming MD/PhD student interested in Alzheimer’s disease, schizophrenia, and the use of omics approaches to better understand disease states. He completed his BSc in Neuroscience at McGill University with an Honours thesis centered on how astrocytes spatially associate with engram neurons during fear memory formation.
Margarita Kapustina (Healthy Cognition Working Group) CV | margokap@student.ubc.ca
PhD Candidate | NSERC Postgraduate Scholar - Doctoral | CIHR Canada Graduate Scholar - Master’s | UBC Four-Year Fellow | UBC Killam Doctoral Fellow
Margarita's research focuses on identifying functionally distinct neuronal subpopulations in layer 6b of the mouse and human neocortex. She completed a BSc in biology at UBC, conducting her undergraduate research thesis and NSERC USRA in our lab, identifying layer 6b neuronal subtypes through big data analyses and in situ techniques.
Julia Groening (Alzheimer’s Working Group) CV | thejuliagroening@gmail.com
Experimental Medicine MSc student | Canada Graduate Research Scholar
Julia is a Master’s student interesting in the cellular mechanisms that underlie Alzheimer’s disease. Julia completed her BSc in Neuroscience at the University of British Columbia. Julia conducted her thesis and other research projects in the MacVicar Lab where she studied the impact of oxidative stress on pericyte and astrocyte dysfunction.
Thomas van der Sloot (Alzheimer’s Working Group) CV
Neuroscience Co-op Undergraduate Student | Wesbrook Scholar | HSBC Emerging Leader Scholar | Thelma Sharp Cook Scholar | NSERC USRA Awardee
Thomas is a fourth-year neuroscience undergraduate student with an interest in neurodegenerative diseases. He is currently serving as the laboratory manager, and will be conducting research looking into the underlying mechanisms behind Alzheimer’s disease by using advanced techniques to investigate the role of key molecules at the sub-cellular level.
Richard Jiang (Epilepsy Working Group)
Cellular and Physiological Sciences Honours Thesis Student
Richard is a fourth-year student in Cellular, Anatomical, and Physiological Sciences. With experience working in a prion lab, he has developed a keen interest in brain morphology and various neurological pathologies. Richard aims to leverage his background in handling large data structures to gain deeper insights into brain function.
Muskan Poddar (Healthy Cognition Working Group) CV
Neuroscience Undergraduate Student | SURE awardee
Muskan is a third-year Neuroscience undergraduate student at UBC, and is interested in exploring cellular mechanisms underlying neurodegenerative disease. At the Cembrowski Lab, she is using computational techniques to investigate changes in cognitive regions of the brain.
Hope Luo (Healthy Cognition Working Group) CV
Cellular and Physiological Sciences Undergraduate Student
Hope is a third-year student in Cellular, Anatomical, and Physiological Sciences. She is passionate about using transcriptomic data to characterize inhibitory interneuron populations in the subiculum. Additionally, she is also developing a web portal to present the lab's progress in a clear and accessible manner for the research community.
Cindy Zhang (Healthy Cognition Working Group) CV
Neuroscience Undergraduate Student | EnSURE Awardee | SBME Synergy Awardee
Cindy is a fourth-year undergraduate neuroscience student at UBC. She is interested in the cellular and molecular mechanisms underlying brain function and dysfunction. Her work will focus on characterizing inhibitory interneuron gene-expression profiles in the hippocampal subiculum using single-cell spatial transcriptomics, with the goal of better understanding how cellular identity contributes to neural circuit function.
Ryan Shen (Healthy Cognition Working Group)
Directed Studies Undergraduate Student (starting Sept 2026)
Ryan is a fourth-year undergraduate student in Behavioural and Cognitive Neuroscience at the University of British Columbia. He is interested in how hippocampal circuits encode and modulate the emotional content of experience. In the Cembrowski lab, Ryan will use histological and microscopy techniques to process and analyze brain tissue from behavioural experiments, with hopes to explore opportunities to integrate computational approaches for quantitative imaging analysis.
Collaborators
Mypinder Sekhon (MD, PhD)
Head, UBC Division of Critical Care Medicine | Intensivist, Vancouver General Hospital
We collaborate to understand the cellular-molecular reorganization of the human brain following traumatic brain injury, hypoxic-ischemia brain injury, and other neurological conditions. Our collaboration is funded by the VGH-UBC Hospital Foundation.
Veronica Hirsch-Reinshagen (MD, PhD)
Neuropathologist, Vancouver General Hospital
We collaborate to identify clinical-pathological correlations in brain disorder and disease including Alzheimer’s disease and schizophrenia. Our collaboration is funded the Canadian Institutes of Health Research and the New Frontiers in Research Fund.
Gary Redekop (MD, MSc)
Neurosurgeon, Vancouver General Hospital
We collaborate to maintain viability of brain tissue removed from epilepsy, and use this tissue to understand seizure-like initiation and propagation in the living human brain. Our collaboration is funded by the Canadian Institutes of Health Research and the New Frontiers in Research Fund.
Mostafa Fatehi-Hassanabad (MD, MSc)
Neurosurgeon, Vancouver General Hospital
We collaborate to maintain viability of brain tissue removed from epilepsy and brain tumors, and use this tissue to understand cellular-molecular function of the living human brain.. Our collaboration is funded by the Brain Canada Foundation, the Canadian Institutes of Health Research and the New Frontiers in Research Fund.
John Maguire (MD)
Neuropathologist, Vancouver General Hospital
We collaborate to identify neuropathological properties of epilepsy. Our collaboration is funded by the Canadian Institutes of Health Research and the New Frontiers in Research Fund.
Ann Marie Craig (PhD)
Professor, University of British Columbia
We collaborate to understand the synaptic reorganization of human brain tissue in epilepsy and Alzheimer’s disease. Our collaboration is funded by CLEAR Alzheimer’s.
Cheryl Wellington (PhD)
Professor, University of British Columbia
We collaborate to understand the cellular-molecular reorganization of the brain following traumatic brain injury and hypoxic-ischemic brain injury. Our collaboration is funded by the USA Department of Defense.
Alumni
Hans Bae: Undergraduate student (now: UBC undergraduate)
Milena Baldauf: Visiting MSc student (now: University Bremen graduate student)
Willis Cao: Honours Thesis undergraduate student (now: UBC undergraduate)
McKinley Carbert: undergraduate student
Raja Choudhary: NSERC USRA undergraduate student (now: UBC Medicine student)
Jaime Conibear: Directed Studies undergraduate student (now: UBC undergraduate student)
William Daniels: Directed Studies undergraduate student (now: UBC Medicine student
Kate Dunne: NSERC USRA and Honours Thesis student (now: NYU graduate student)
Madeline Elder: Honours Physics undergraduate student (now: UBC Medicine student)
Sarah Erwin: Technician and MSc student
Riya Gandhi: SBME Synergy Co-op student (now: Master’s of Global Health student)
Axel Guskjolen: NSERC Postdoctoral Researcher (now: editor, Neuron)
Aahana Kanyal: SBME co-op student (now: Tufts University graduate student)
Rennie Kendrick: Fulbright Scholar (now: Stanford University graduate student)
Adrienne Kinman: NSERC CGS-D PhD student (now: Princeton University postdoctoral researcher)
Esther Kim: Undergraduate student (now: UBC undergraduate)
Timothy O’Leary: Research assistant (now: Research Assistant, Bamji lab)
Nadine Plett: Directed Studies undergraduate student (now: UBC undergraduate student)
Aaryan Rampal: Computer Science Directed Studies Undergraduate Student (now: industry)
Cat Schwab: undergraduate student
Kaitlin Sullivan: NSERC CGS-D PhD student (now: Princeton University CIHR postdoctoral researcher)
Adi Swaro: Wesbrook Scholar undergraduate student (now: UBC MD student)
Ali Tarik: Directed Studies and co-op undergraduate student (now: UBC undergraduate student)
Joshua Tindall: Honours Physics undergraduate student (now: McGill graduate student)
Jennifer Tsai: Biotalent Canada - Waterloo co-op student (now: Waterloo undergraduate student)
Lucie Wang: Directed Studies and co-op undergraduate student (now: UBC undergraduate student)
Stacy Wang: Undergraduate student (now: UBC undergraduate)
Sydney Wood: Visiting MSc student (now: University of Nottingham graduate student)
Angela Zhang: NSERC USRA undergraduate student (now: UBC Medicine student)
Ming Zhang: SURE undergraduate student
Mia Zheng: Biology Honours Thesis Undergraduate Student (now: McGill graduate student)
Lauren Zung: Directed Studies undergraduate student (now: UBC Data Science MSc student)
Publications
See also Google Scholar and Pubmed
Kinman, A.I.*, Kraus, L.*, Cembrowski, M.S.. The subiculum: cell-type-specific composition, computation, and function. Trends in Neurosciences 2026.
Futhey, N.C.*, Vila-Rodriguez, F., Stockmanski, S., Gregory, E., Honer, W.G., Arranz, B., Ramos, B., Escanilla, A., Vera-Montecinos, A., Cembrowski, M.S.*, Hirsch-Reinshagen, V.* Integrated clinical and postmortem profiling in schizophrenia reveals a cognitive subtype linked to cerebrovascular disease. Translational Psychiatry 2026.
Campbell, R.E.*, Zhang, M.Y.#, Merryweather, D.*, Cembrowski, M.S. Divergent hippocampal output via covariant local and long-range neuronal structure. Progress in Neurobiology 2026
Bird, J.D., Cembrowski, M.S. Sekhon, M.S, Omics-based Approaches for Acute Brain Injury Research. Current Opinion in Critical Care 2026
Kapustina, M.*, Bristow, B.N.*, Cembrowski, M.S. Distinct Layer 6b transcriptomic subtypes parcellate the cortical mantle. Progress in Neurobiology 2025.
Sullivan, K.E., Kapustina, M.*, Bristow, B.N.*, Cembrowski, M.S. Quantification and analysis of multiplexed fluorescence in situ hybridization data using open-source tools. STAR Protocols 2025.
Swaro, A.*, Bristow, B.N.*, Anwer, M.*, Zhang, A.A., Kraus, L., Erwin, S.R., Stach, T.R., Sullivan, K.E., Gandhi, R., Fan, J., Cheng, W.H., Wellington, C.L., Cembrowski, M.S. Widespread and cell-type-specific transcriptomic reorganization following mild traumatic brain injury. Cell Reports 2025.
Hammond, B.P., Zia, S., Hahn, E., Kapustina, M., Lange, T., Friesen, S., Manek, R., Lee, K.V., Castellanos-Molina, A., Bretheau, F., Cembrowski, M.S., Kerr, B.S., Lacroix, S., Plemel, J.R. CSF-1R ligands promote microglial proliferation but are not the sole regulators of developmental microglial proliferation. Development 2025.
Kinman, A.I., Merryweather, D.N., Erwin, S.R., Campbell, R.E., Sullivan, K.E., Kraus, L., Kapustina, M., Bristow, B.N., Zhang, M.Y., Elder, M.W., Wood, S.C., Tarik, A., Kim, E., Tindall, J., Daniels, W., Anwer, M., Guo, C., Cembrowski, M.S. Atypical hippocampal excitatory neurons express and govern object memory. Nature Communications 2025.
Kapustina, M.*, Zhang, A.A.*, Tsai, J.Y.J., Bristow B.N., Kraus, L., Sullivan, K.E., Erwin, S.R., Wang, L., Stach, T.R., Clements, J., Lemire, A.L., Cembrowski, M.S. The cell-type-specific spatial organization of the anterior thalamic nuclei of the mouse brain. Cell Reports 2024.
Guskjolen, A., Cembrowski, M.S. Engram neurons: Encoding, consolidation, retrieval, and forgetting of memory. Molecular Psychiatry 2023.
Sullivan, K.E., Kraus, L., Kapustina, M., Wang, L., Stach, T., Clements, J., Lemire, A., Cembrowski, M.S. Sharp cell-type-identity changes differentiate the retrosplenial cortex from the neocortex. Cell Reports 2023.
O’Leary, T.P.*, Kendrick, R.M.*, Bristow, B.N., Sullivan, K.E., Wang, L., Clements, J., Lemire, A., Cembrowski, M.S. Neuronal cell types, projections, and spatial organization of the central amygdala. iScience 2022.
Erwin, S.E.*, Bristow, B.N.*, Kendrick, R.M., Sullivan, K.E., Marriot, B., Wang, L., Clements, J., Lemire, A., Jackson, J., Cembrowski, M.S. Spatially patterned excitatory neuron subtypes and projections of the claustrum. eLife 10:e68967. 2021.
Sullivan, K.E.*, Kendrick, R.K.*, Cembrowski, M.S. Elucidating memory in the brain via single‐cell transcriptomics. Journal of Neurochemistry 2021.
O’Leary, T.P.*, Sullivan, K.E.*, Wang, L., Clements, J., Lemire, A., Cembrowski, M.S. Extensive and spatially variable within-cell-type heterogeneity across the basolateral amygdala. eLife 9:e59003. 2020.
eLife “Striking Image”
Erwin, S.R.*, Sun, W.*, Copeland, M., Lindo, S., Spruston, N., Cembrowski, M.S. A sparse, spatially biased subtype of mature granule cell dominates activity in hippocampal-associated behaviors. Cell Reports 31(4):1-2. 2020..
Cembrowski, M.S. Single-cell transcriptomics as a framework and roadmap for understanding the brain. Journal of Neuroscience Methods. 326:1-7. 2019.
Cembrowski, M.S., Spruston, N. Heterogeneity within classical cell types is the rule: lessons from hippocampal pyramidal neurons. Nature Reviews Neuroscience, 20(4):193-204. 2019.
Recommendation on Faculty of 1000.
Cembrowski, M.S., Wang, L., Lemire, A., DiLisio, S.F., Copeland, M., Clements, J., Spruston, N. The subiculum is a patchwork of discrete subregions. eLife 7, doi:10.7554/eLife.37701, 2018.
Research Highlight. Lewis, S. Patchwork subiculum. Nature Reviews Neuroscience, 20(1):3. 2019.
Recommendation on Faculty of 1000.
Cembrowski, M.S., Phillips, M.G., DiLisio, S.F., Shields, B.C., Winnubst, J., Chandrashekar, J., Bas, E., Spruston, N. Dissociable structural and functional hippocampal outputs via distinct subiculum cell classes. Cell 173(5): 1280–1292, 2018.
Research Highlight. Whalley, K. A regional divide. Nature Reviews Neuroscience, 19(7):390. 2018.
Cembrowski M.S., Menon, V. Continuous variation within cell types of the nervous system. Trends in Neurosciences 41(6): 339-350, 2018.
Bloss, E.B., Cembrowski, M.S., Karsh, B., Colonell, J., Fetter, R.D., Spruston, N. Single excitatory axons form clustered synapses onto CA1 pyramidal cell dendrites. Nature Neuroscience 21(3): 353-363, 2018.
Cembrowski, M.S., Spruston, N. Integrating results across methodologies is essential for producing robust neuronal taxonomies. Neuron 94(1): 747-751, 2017.
Cembrowski, M.S., Wang., L., Sugino, K., Shields, B.C., Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, 10.7554/eLife.14997, 2016.
Bloss, E.B., Cembrowski, M.S., Karsh, B., Colonell, J., Fetter, R., Spruston, N. Structured patterns of dendritic inhibition support branch-specific forms of integration in CA1 pyramidal cells. Neuron 89(5): 1016-1030, 2016.
Cembrowski, M.S., Bachman, J.L., Wang, L., Sugino, K., Shields, B.C., Spruston, N. Spatial gene-expression gradients underlie prominent heterogeneity of CA1 pyramidal neurons. Neuron 89(2): 351-368, 2016.
·Featured article of the issue. Previewed by Tushev, G. and Schuman, E.M. Rethinking Functional Segregation: Gradients of Gene Expression in Area CA1. Neuron 89(2):242-243, 2016.
·Of Outstanding Interest. Mallory, C.S. and Giocomo, L.M. Heterogeneity within hippocampal place coding. Review, Current Opinion in Neurobiology 49:158-167, 2018.
·Highlighted reference (1 of 6). Soltesz, I. and Losonczy, A. CA1 pyramidal cell diversity enabling parallel information processing in the hippocampus. Review, Nature Neuroscience 21(18): 484-493, 2018.
Of Special Interest. Valero, M. and de la Prida, L,M. The hippocampus in depth: a sublayer-specific perspective of entorhinal–hippocampal function. Review, Current Opinion in Neurobiology 52:107-114, 2018.
Of Special Interest. Suvrathan, A. Beyond STDP – Towards Diverse and Functionally Relevant Plasticity Rules. Review, Current Opinion in Neurobiology 54:12-19, 2019.
Kim, Y.*, Hsu, C.-L.*, Cembrowski, M.S., Mensh, B.D., Spruston, N. Dendritic sodium spikes are required for long-term potentiation at distal synapses on hippocampal pyramidal neurons. eLife 4, doi:10.7554/eLife.06414, 2015. *: authors contributed equally
·Recommendation on Faculty of 1000.
Choi, H., Lei, Zhang, L., Cembrowski, M.S., Sabottke, C.F., Markowitz, A.L., Butts, D.A., Kath, W.L., Singer, J.H., Riecke, H. Intrinsic bursting of AII amacrine cells underlies oscillations in the rd1 mouse retina. Journal of Neurophysiology 112(6): 1491-1504, 2014.
Ke, J., Wang, Y., Borghuis, B.G., Cembrowski, M.S., Riecke, H., Kath, W.L., Demb, J.B., Singer, J.H. Adaptation to background light enables contrast coding at rod bipolar cell synapses. Neuron 81(2): 388-401, 2014.
·Recommendation on Faculty of 1000.
Cembrowski, M.S., Logan, S., Tian, M., Jia, L., Li, W., Kath, W.L., Riecke, H., Singer, J.H. The mechanisms of repetitive spike generation in an axonless retinal interneuron. Cell Reports 1(2): 155-166, 2012.
Interested in joining?
Who we are
We are a multidisciplinary group integrating a mix of neuroscience, engineering, physics, math, and computation. Our lab members span these academic backgrounds, and work in a collaborative, friendly, and exciting atmosphere.
What we look for
Applicants with backgrounds in neuroscience, engineering, physics, math, and computer science are welcome, and underrepresented minorities are strongly encouraged to apply. We seek people that are passionate about science and interested in working in a collaborative, dynamic, and supportive lab environment.
How to apply?
All applicants: Applicants interested in the Cembrowski Lab should contact Mark via email at mark.cembrowski [ at ] ubc.ca with a CV, unofficial academic transcript, and a brief and specific explanation of why they are interested in the lab. Emails that do not meet these criteria, or that do not address following additional points, are not responded to.
Postdoctoral applicants: A brief cover letter (2-3 paragraphs) summarizing past work and future goals should be included in the email.
MSc/PhD applicants: Graduate school applicants can apply to the lab through programs in Neuroscience, Biomedical Engineering, Bioinformatics, Experimental Medicine, or Mathematics. In all of these programs applicants are directly paired with a supervisor (i.e., no rotations), and thus applicants should reach out to Mark before applying to assess fit and availability. Grad school applicants are recommended to check out our Frequently Asked Questions, and are also recommended to these great general tips for applying to grad school. Applications for a September 2026 start date are closed. September 2027 is the earliest next start date in consideration, and interested applicants should reach out in Fall 2026.
MD-PhD applicants: MD-PhD applicants apply concurrently through the UBC MD and MD-PhD program, and should follow the instructions above for PhD applicants to have Mark as their potential PhD supervisor. Applicants should include their MCAT scores in their initial email, and review the UBC MD-PhD Frequently Asked Questions.
Technicians: A brief explanation of research experience and previously used techniques should be included in the email.
Undergraduate students: We do not take students for volunteer positions, as this perpetuates existing inequities in science. Undergraduate students begin in the lab via credit-based opportunities (e.g., via Directed Studies, Honours Thesis), and have at least a third-year standing when joining. Summer positions and other full-time positions (e.g., Co-op) are prioritzed for students that have already done credit-based work within the lab. Note: our undergraduate positions are currently full, but students are encouraged to reach out to see if there may be upcoming space. Due to high demand, undergraduate positions are typically booked 8-12 months in advance.
Media
Interviews and presentations: some highlights of our research
Mark interviewed in Nature Methods
Mark presents research at the Allen Institute Showcase Symposium
Gallery: A collection of our striking images from the hippocampus and beyond. Our images have been featured in science + art series like the Cell Picture Show and NeuroArt.
The activated dentate gyrus
Granule cells of the dentate gyrus (red), induced to drive activity (green: activity reporting via cFos immunohistochemistry).
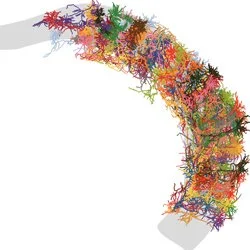
Axonal tapestry
Axons from two different coloured subiculum projections interleave and course towards the thalamus.
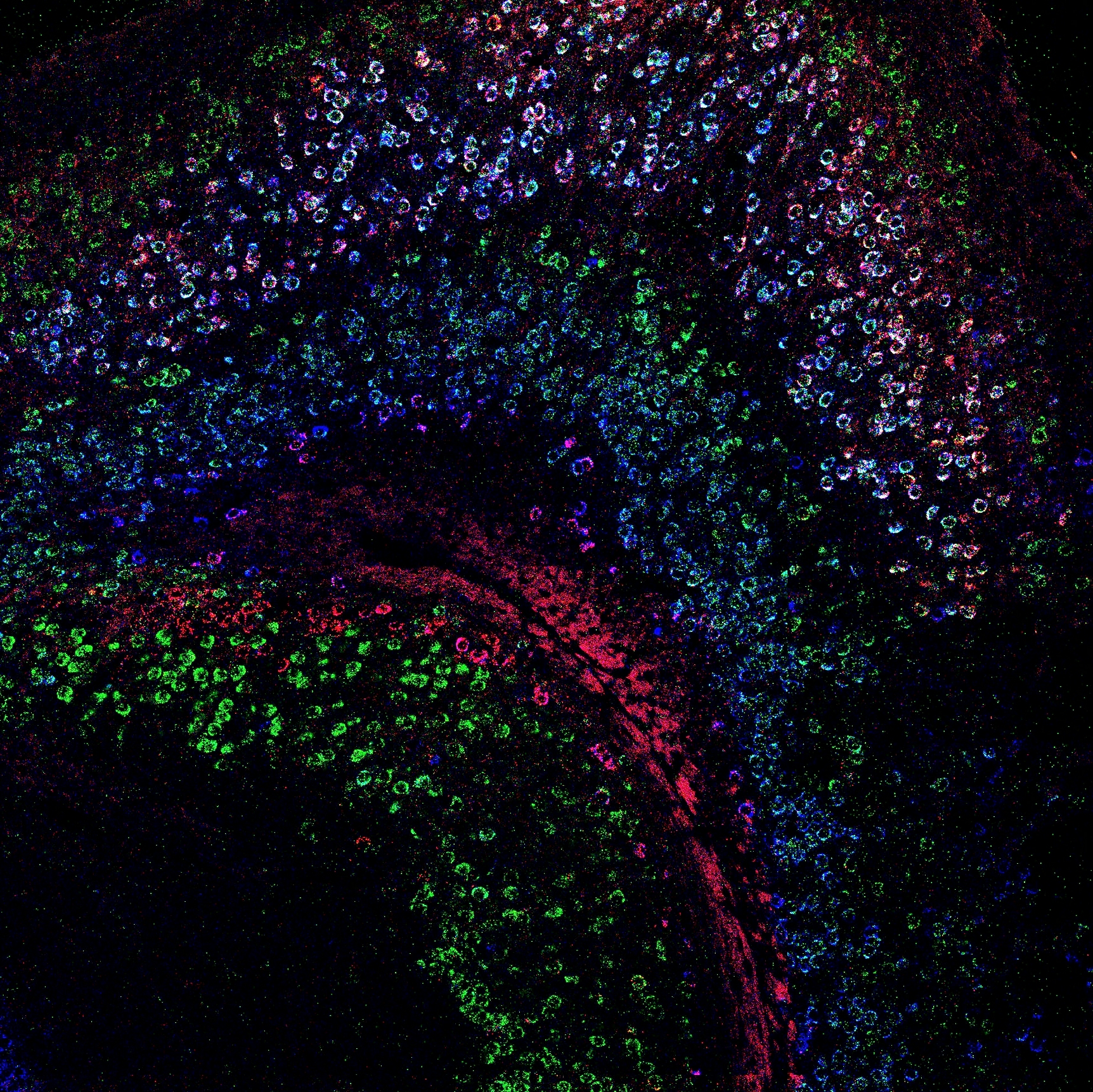
Neuronal pointillism
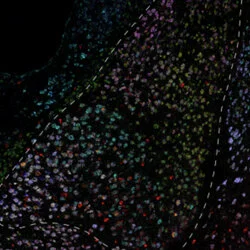
Overview of the hippocampus and entorhinal cortices, with Dkk3, Ly6g6e, and Pcp4 labeled via in situ hybridization
A forest of mossy cells
Mossy cells of the hilus received their name due to their elaborate moss-like dendritic elaborations.
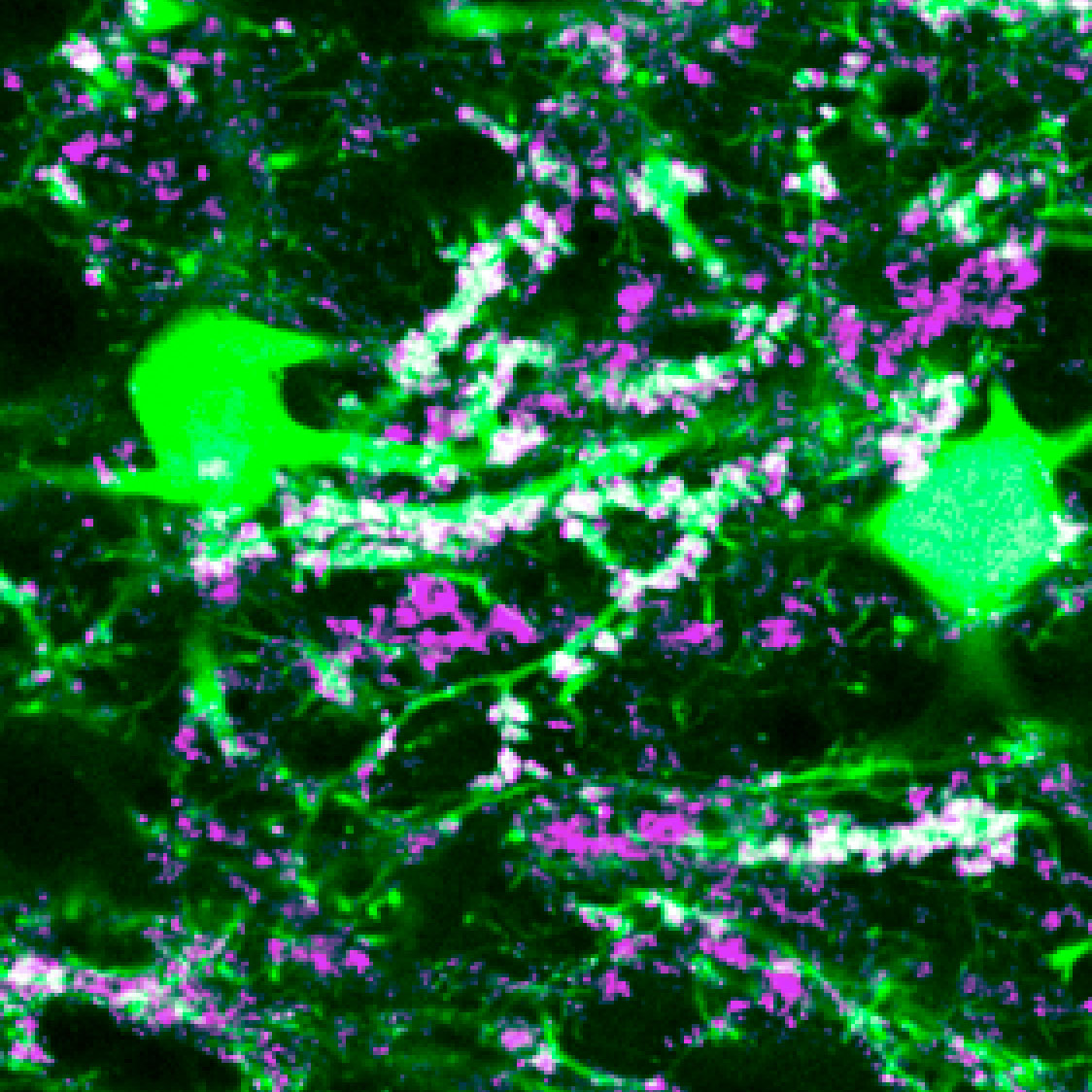
Flowering mossy branches
Mossy cell branches are decorated with inputs from large glutamatergic terminals (via Vglut1 immunohistochemistry).
The coloured hippocampus
The major cell types of the hippocampus (blue: granule cells; red/yellow/green: CA3/CA2/CA1 pyramidal cells).
Funding and support
We are grateful to the following groups for supporting our research.